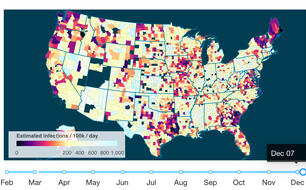
Several members of the Yale Public Health Modeling Unit have developed and implemented a statistical model which uses reported time-series of COVID-19 diagnoses and deaths to estimate the current number of individuals infected with COVID-19, the cumulative incidence of infection, and the effective reproduction number at the county- and state-levels in the United States.
Estimates of the current number of local infections, the trajectory of such infections over time, and the fraction of the local community that has already been infected are critical for understanding the impact of interventions such as mask wearing, business closures, and physical distancing. Furthermore, these estimates can help both policymakers and individuals tailor their decisions to local epidemic conditions, which is not possible based on case reports and deaths which are incomplete and lagged indicators of the epidemic.
This Bayesian nowcasting model, which is updated daily and available at covidestim.org, leverages information about disease progression and severity, delays in diagnosis associated with the disease incubation period, care seeking behavior, diagnostic processing times and reporting practices to estimate time series of incident infections which are compatible with observable data.
This modeling tool was developed as an R package and documented in our GitHub repository, and can easily be run for a county or state of interest. However, the ultimate goal of offering nowcasts for every county and state necessitated the ability to execute more than 3000 runs of the model - about eight months’ worth of computation - every day. YCRC was instrumental in discussing and exploring several options across different vendors and computing platforms for achieving this challenging goal. The resulting system has proven to be cost-effective, highly reliable, and easy to manage, demonstrating the feasibility of quickly creating large-scale public-health infrastructure to respond to emergent infectious diseases.